 trj was done by gmx trjconv, but I don't think this is possible anymore (see gmx trjconv supported trajectory formats in GROMACS 5 vs GROMACS 2021). In recent versions of GROMACS trj files are no longer supported, probably as they were not in a binary format, hence took too much disk space (see Trajectory files formats in GROMACS 2021 documentation). This is also why it tells you the header is wrong. 
trj files were used in "old" versions of GROMACS to store trajectories (see trj file format documentation of GROMACS 5), and this is what VMD is awaiting, not the output of gmx dump. Trj files are NOT the output of gmx dump. 
You can use this page to link to that part and include additional information about your contribution.I don't think that the output of gmx dump can be read by VMD: the goal of this command is to provide a human-readable output, not to convert one trajectory format into another. If you are making a contribution by adding information to an existing Part or creating a new Part, you must document your contribution on the Part's Main Page on the Registry for your team to be eligible for this criteria. Make sure to have a video player installed that supports your chosen format, and also determine a good folder to place your video in.īelow is a sample animation of P0DTR5 and below that is P0DTR4. When you have finished making your choices, click make movie, and your animation is done. From the Movie Generator window, the movie options can be tweaked to determine the axis of rotation, and the duration of the video can be changed to fit the need. To make a video of the protein, find the Extensions tab, scroll down to Visualization, and click on Movie Maker on the drop down menu. Once you click apply, your molecule will be displayed in the OpenGL Display tab, and can be manipulated from there to observe protein features. An easy, informative, and aesthetic choice would be the secondary structure coloring, and NewCartoon drawing method, but there are more options to explore for the given modelling need. A new window will appear titled “Graphical Representations” which will give you a litany of options for coloring, drawing types, and other material appearance. To display the molecule, click on the graphics tab, and click on representations. This should then pop up in the VMD Main window, and display your “molecule”, the number of atoms and other information. Once VMD and your desired protein pdb file is downloaded, the protein can be observed by first clicking on the file tab on the VMD Main window, then find your pdb file and click load. 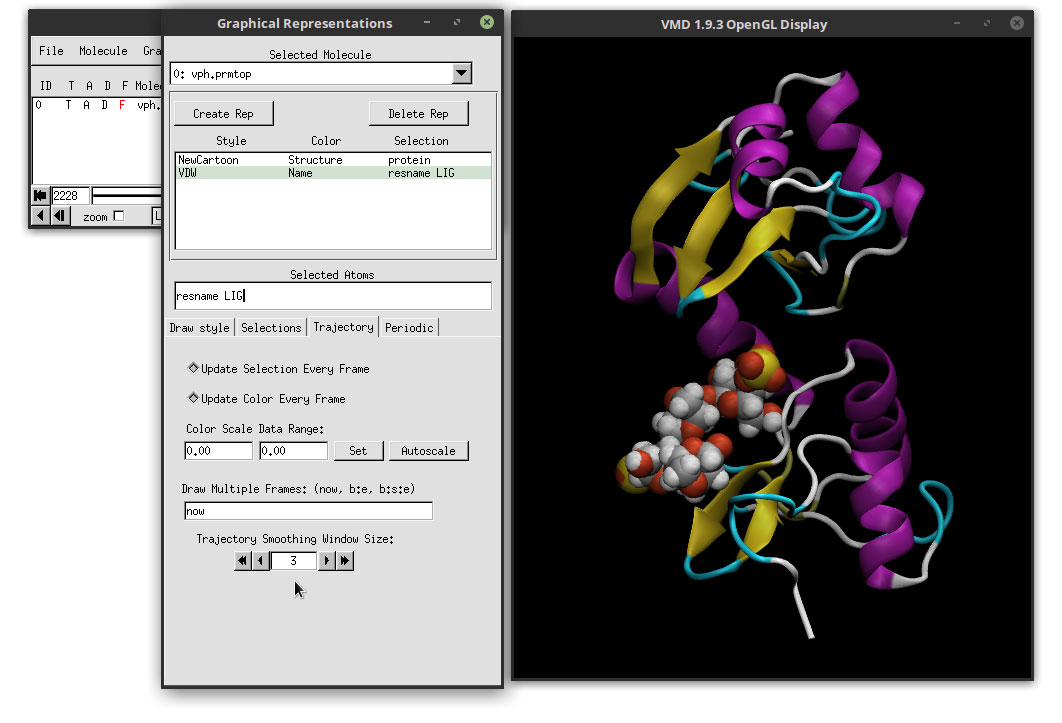
VMD will be downloaded from the internet, and thus, will be unrecognized by Windows at first- make sure foreign programs outside Microsoft store are allowed under your default settings. Otherwise, the file will likely be a gz file, of which, VMD cannot do anything with. 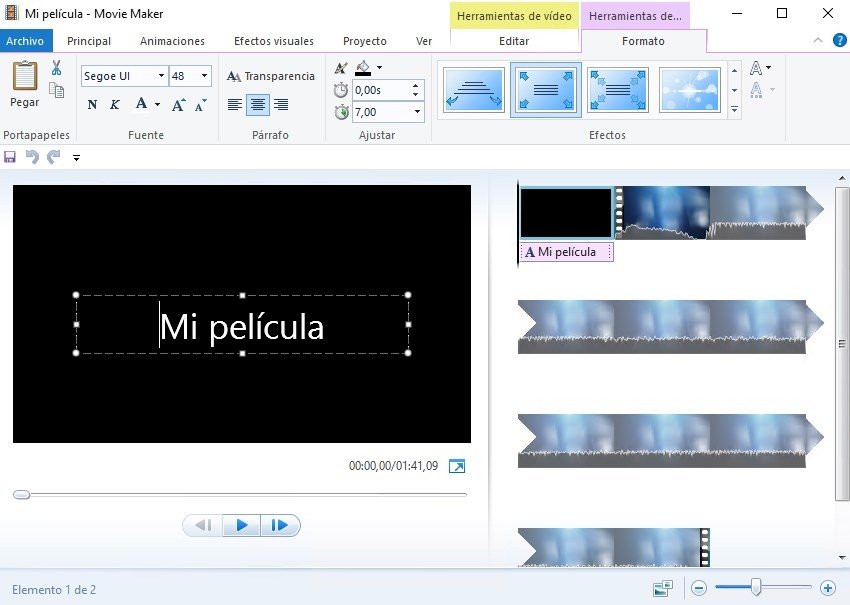
When downloading the pdb file, make sure to click on the top right download file option, and scroll down to the pdb format option. Go to the RCSB site and search for your protein's PDB code, click on the doi link, and download the structure coordinates. Under the "Structures" section, there will be a PDB code for your protein under the model. Go to the UniProt site and search for your protein's name. Once the program is downloaded, open it and now you are ready to find the. Go to this page and make an account to download the appropriate version of VMD for your OS. For those working with proteins, this is a guide on how to use VMD to make models of those proteins.įirst, you need to download VMD. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
